This piece was written live during the 5th annual RNA Symposium: Processing RNA. Follow us on Twitter or the tag #umichrna
Live Blogger: Emily Glass
Editor: Zoe Yeoh
Tracy Johnson, Ph.D., a researcher at the University of California – Los Angeles, uses yeast (S. cerevisiae) to study gene regulation and expression with a focus on the spliceosome. The spliceosome is a dynamic cellular machine made up of 5 ribonucleoprotein subunits that is responsible for creating mature messenger RNA (mRNA). During precursor mRNA (pre-mRNA) splicing, the spliceosome removes the many non-coding sequences (introns) that eukaryotic DNA produces in protein-encoding genes during transcription and splices together the coding sequences (exons) to allow for mature mRNA production.
Dr. Johnson begins her talk by getting listeners to think about the first steps in studying a complicated machine, like the spliceosome. She posits that you would likely start taking parts away from the machine to see how each piece affects its function. But, just as you would rather figure out how a car works by experimenting on a basic model instead of a Ferrari, she uses yeast as her model organism to study spliceosome function in eukaryotes instead of studying more complex mammalian models.
A series of clever experiments done by Dr. Johnson’s lab has since teased out specific interactions between transcription machinery and the spliceosome. Recent advances in the field had shown that post-translational modifications such as methylation of the histone H3K36 could regulate exon inclusion in mature mRNA, but there were still some unanswered questions about other effects this methylation may have on splicing. Building on this finding, Dr. Johnson’s lab has shown that the trimethylation of the histone H3K36 by the histone methyltransferase Set2 influences pre-mRNA splicing accuracy on a much broader scale. Yeast with both a H3K36 point mutation and Set2 deletion were unable to splice together a variety of proteins. After this finding, they came up with the hypothesis that H3K36 methylation actively recruits the spliceosome.
In considering how H3K36 methylation could recruit the spliceosome, Dr. Johnson began to consider the different proteins that had been shown to bind to H3K36 and targeted Eaf3, a chromodomain protein. Sure enough, yeast Eaf3 knockouts showed the same unspliced protein as the H3K36 and Set2 loss-of-function mutations. Furthermore, in an exciting finding, different spliceosome machinery was found to co-precipitate with Eaf3, indicating a direct interaction between these RNA’s.
This led them to the question: how does Eaf3 interact with the spliceosome? Previous work had shown that Eaf3 interacts with the essential splicing factor Prp45, so Dr. Johnson decided to start looking there. She found that Prp45 levels decrease in yeast with Eaf3 knocked out, an important discovery that indicates Eaf3 is essential for the recruitment of Prp45 and splicing accuracy.
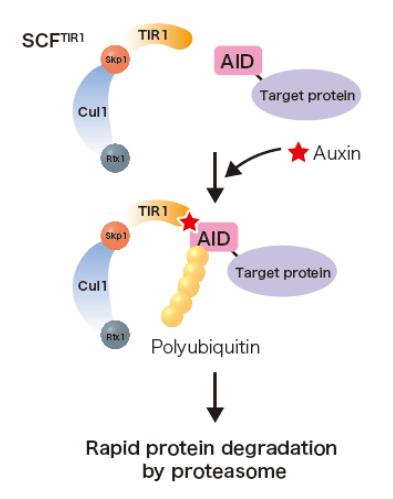
Even though all of the discussed work thus far was groundbreaking by itself in the field of gene expression, Dr. Johnson wanted to further investigate the importance of Prp45. As Prp45 is an essential gene, researchers in Dr. Johnson’s lab couldn’t knock out Prp45 completely to discern its function as they had done with the other genes of interest. So, they began their work using a truncated version of Prp45 and found that Prp45 is necessary for the methylation of H3K36. This was a novel finding as this suggests a bidirectional regulation of gene expression by transcription machinery and the spliceosome. To further characterize how interdependent these two factors are, Dr. Johnson wanted to deplete Prp45 to see what would happen to H3K36 methylation. As Prp45 is an essential gene, they had to use an Auxin-induced degron system (AID), which allows for the degradation of proteins in vivo.
A completely unexpected finding took place: complete loss of Prp45 does not lead to loss of H3K36 methylation. Instead, it dysregulates it in the opposite direction! Methylation of H3K36 was in excess compared to when Prp45 was present. Dr. Johnson ends her talk by stating that, taken together, these findings suggest that Prp45 regulates H3K36 methylation co-transcriptionally, a novel mechanism by which gene expression is regulated. Her next endeavor is to understand how the cell’s environment may influence this pathway.
Tracy Johnson received her B.A. in Biochemistry and Cell Biology from UCSD, and her Ph.D. in Biochemistry and Molecular Biology from UC Berkeley. She went on to be a Jane Coffin Childs postdoctoral fellow at the California Institute of Technology where she began to study RNA splicing, after which she joined the faculty at UC San Diego. She moved to UCLA in 2013 to join the faculty in Molecular, Cell, and Developmental Biology where she is the Keith and Cecilia Terasaki Presidential Endowed Chair.

