Coming to you LIVE from the 3rd annual RNA Symposium: Advancing RNA Bioscience into Medicine. Follow us on Twitter or the tag #umichrna!
Live blogger: Sarah Kearns. Editor: Whit Froehlich.
Suppose you have some extremely important information in the form of a blueprint and it’s your job to protect it. It’s not just a blueprint for a top-secret location – it’s the blueprint to life; it specifies how every cell in the body should function.
Needless to say, you have to keep this information in a secure spot. This is exactly what eukaryotic cells do with their DNA by storing it in the nucleus, a membrane-enclosed compartment. The genetic material itself never leaves the nucleus; instead it’s transcribed (essentially making a copy through the genetic pairing process) as messenger RNA (mRNA) molecules that leave the nucleus. From there, to be translated into a protein product, mRNA has to go to a molecular machine called the ribosome. This molecular sandwich, also made of RNA, “reads” the information on the mRNA and translates it into the amino acid sequence that it spells out, making proteins.
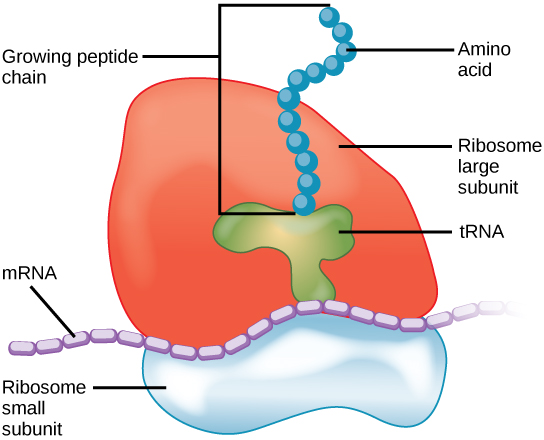
An understanding of how the ribosome works is important for a scientific understanding of life due to its role in protein production. As the RNA instructs the ribosome to assemble amino acids in the proper order, the new polypeptide – or protein – starts to fold into a compact blob. This final blob shape is important because a protein’s function is significantly determined by its structure. As such, if the protein does not fold into the correct shape, then it cannot do what it’s supposed to do in the cell. Many disease states, including sickle cell anemia, Parkinson’s Disease, and Alzheimer’s Disease, are related to protein misfolding.
Dr. Jonathan Weissman, a professor of cellular and molecular pharmacology at the University of California, San Francisco, has helped develop a key technique called ribosome profiling that monitors protein translation. Pairing together this large-scale approach with smaller mechanistic investigations, Dr. Weissman looks at the role of the ribosome in aiding correct protein folding.
The ribosome’s work of translation is itself a very dynamic process. So far, structures obtained from x-ray crystallography and cryo-EM allow for a reconstruction of a simulated video, but the dream would be to actually visualize this process in the cell. A step closer is getting the full time course of translation, which Dr. Weissman has been working on through ribosome profiling.
Ribosome profiling globally profiles protein translation. Unlike typical analysis of a lifetime of behaviors, this technique takes a snapshot of all of the ribosomes within a cell at a particular time. The mRNA sequences that are protected by the ribosome from the processing process are counted, yielding a measurement of the mRNA sequences being translated when the analysis is performed. This technique can further monitor when other proteins are interacting with the ribosome or mRNA.

The precision achieved by combining billions of these footprinting experiments can tell us how much RNA and protein are being synthesized and where along the known gene sequence that transcription is occurring. The most abundant proteins, he finds, have yet to be marked on the genetic code, suggesting a significant number of unidentified open reading frames within the genome.
Ribosome profiling as well more precisely reveals the rate of protein expression, which is currently measured by counting mRNAs. Taking F-ATPase as an example, because there are different subunits in different amounts that create the protein, he finds that the ribosome density along the mRNA correlates with the ratios of the different domains in the final protein. This makes sense, as it results in efficient production of the protein.
The next question is: where is translation occurring? Because the cell is relatively large, at least compared to a single ribosome, to make sure the protein synthesis is occuring in the right place, the mRNA might be brought to the correct spot within the cell before it’s translated.
By adding a tag specifically to the endoplasmic reticulum (ER), they can look at the specific ribosome profile occuring there. Proteins that need to travel to the membrane need to be bundled and sent there through signalling done by the ER. Comparing the ER-ribosome footprints with the overall ribosome footprints, they find that proteins destined for secretion through the membrane, or installation in the membrane, are translated at the ER. So this tag works to identify known and localized mRNA sequences.

To test the same technique with other organelles, they add the tag to mitochondria instead of the ER, but run into some complications (what would science be without its annoying barriers?). This is because there is a protein complex (ERMES) that tethers the mitochondria to the ER. Nevertheless, they were able to tease apart the ribosome profiles by enriching the cell for either mitochondria or ER. Adding this tag to other organelles, proteins, and other factors will allow discovery of the spatiotemporal map of translation across the whole cell!
But the ER has a tough job! Not only is it responsible for trafficking membrane proteins, it has roles in protein synthesis and stabilization as well. With proteins having variable lengths, amino acid compositions, and electrochemical properties, there are many different routes in protein trafficking and protein folding, each of which can malfunction. The resulting misfolded or misplaced proteins can causes diseases including cystic fibrosis, hypercholesterolemia, and many others.
Chaperone proteins help with both of these problems (localization and folding). By interacting with misfolded proteins, they pull on tangled polypeptide strands, not unlike how we untangle pocketed headphones, to get the protein into a correct conformation. Because there are so many different types of proteins, chaperones are localized at the final destination of the protein. As such, membrane protein chaperones, like EMC, are at the membrane surface.
By using the tagging technology previously discussed, Dr. Weissman and his team added a tag onto the EMC to find out what ribosomal substrates were being synthesised. It turns out there’s a sequence-specificity for transmembrane domains and hydrophobic patches. These interactions of the EMC and the ribosome occur during translation to ensure that there are chaperones at the site of protein synthesis, avoiding any downtime prior to its proper folding. This suggests that the ribosome, too, has to be localized.
By putting together the results from ribosome profiling and tagging, a map of where transcription occurs is made, raising questions of how these multisubunit complexes are being transported in the crowded cell.
