Live Blogger: Emily Eberhardt
Editor: Christian Greenhill
This piece was written live during the 6th annual RNA Symposium: Towards our Future of RNA Therapeutics, hosted by the University of Michigan’s Center for RNA Biomedicine. Follow MiSciWriter’s coverage of this event on Twitter with the hashtag #umichrna.
Dr. Wendy Gilbert’s lab Twitter biography is simple: We love RNA. However, the intricate details of mRNA specific regulation are complex and tightly-regulated. Dr. Wendy Gilbert starts her presentation with a bold image of a female superhero with the title “Control.” It is immediately clear that she is passionate about her lab’s research and seeks to understand the control of mRNA regulation in the process of translation. Dr. Gilbert has good reason to be interested in better understanding this process, as misregulation has dire consequences: heritable diseases and cancer.
Why is translation initiation an important process?
Dr. Gilbert emphasizes that regulation of gene translation is fundamental to both the small scale of individual cells and on the global scale, in relation to the entire human body. As Dr. Gilbert states, “we know some mRNAs are inefficiently translated while others are robustly translated, and this varies widely.” She then states that translation initiation is rate-limiting, as it requires the assembly of initiation factors and ribosomes, where the small subunit of the ribosome needs to be recruited to mRNA, scan for the start codon, and recruit the large subunit before translation can proceed. This process results in the production of a vast array of proteins in specific regions of the cell, in response to real time needs of the cell.
What is a 5’ UTR?
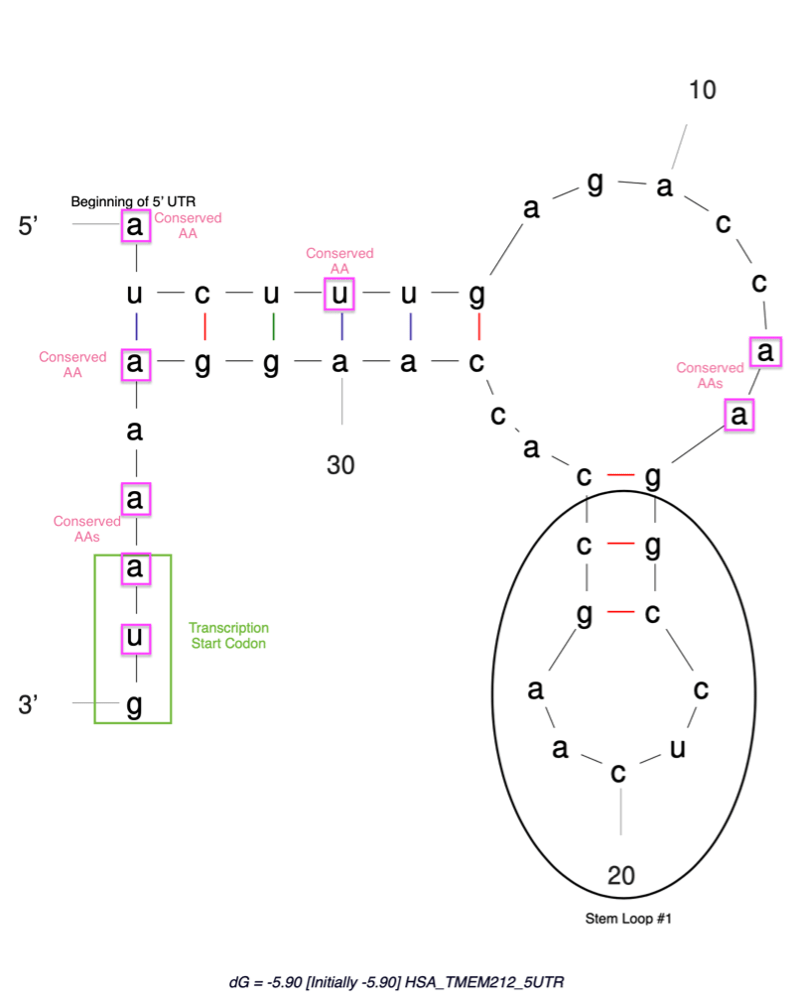
Directly upstream of the start codon of a gene, the five prime untranslated region (5’ UTR) of messenger RNA (mRNA) contains rich elements that affect gene expression (Figure 1). As described above, these regions affect the recruitment of ribosomes to the start codon of mRNA and affect both the translation efficiency and the composition of the cellular proteome. These features have recently gained attention, with researchers such as Dr. Wendy Gilbert expressing interest in the features of 5’ UTRs that coordinate genome-wide translational control. Dr. Gilbert states that the 5’ UTR region is responsible for orders of magnitude in differences in ribosomal recruitment, and thus, gene expression.
How are 5’ UTRs studied?
5’ UTRs are studied using high-throughput methods. These methods allow for the testing of large numbers of biological materials for binding to targets, such as ribosomal binding to mRNAs. Generally, these methods have left a gap in understanding between the quantitative values generated and the differences visualized in translation activity of specific mRNAs. Therefore, Dr. Gilbert and her lab developed a new method of screening these regions to determine how individual regulatory features guide and control translation, called direct analysis of ribosome targeting (DART).
What is DART and how is it used to study 5’ UTRs?

Recently detailed in a publication by Dr. Gilbert’s Lab, DART is used to characterize thousands of 5’ UTRs concurrently. This method relies on a DNA oligonucleotide library containing over 12,000 sequences that correspond to full-length 5’ UTR sequences that flank over 4,000 genes (Figure 2). These sequences were specifically chosen by Dr. Gilbert and her team to focus on the identification of functional elements within the 5’ UTR. As Dr. Gilbert explains, “each initiation factor binds to nucleotides in the 5’ initiation region. The textbook view is that the 5’ UTR is just a landing pad for initiation factors, as long as it is relatively unfolded. However, this is unlikely due to DNA sequence preference and a dependency on specific 5’ UTRs and their interaction with initiation factors.” She continues by stating that the library of sequences can be incubated with initiation factors and ribosomes, before being separated using centrifugation, and then analyzed to determine what sequences are specifically bound to ribosomes and thus actively being translated into protein products. This can then be used to determine which 5’ UTRs inhibit or promote translation.
Dr. Gilbert then outlines how DART has been used by her lab to determine how specific elements within the 5’ UTR prevent 5’ UTR scanning and subsequent protein synthesis, such as stable, secondary structures and C-rich motifs in the mRNA. Other features, such as oligo-uridine regions, promote translation. These features can be characterized and given a “ribosome recruitment score,” which correlates to their effects on gene expression.
What’s next?
Dr. Gilbert’s team intends to continue investigation into the modular nature of mRNA to determine how the features of the 5’ UTR work together to recruit initiation factors. As she concludes her presentation, she emphasizes that the “5’ UTR is an enormous effector of protein output” and implies that her future work will not only focus on utilizing tools such as DART, but also on combining the understanding of these features to generate therapeutic mRNAs that could supplement protein in disease.
Why should I care?
As researchers continue to better understand the roles of 5’ UTR in cellular proteome composition, correlations between the quantitative values generated by high-throughput methods and the direct effect on normal and pathological disease regulation can be made. The method presented by Dr. Gilbert today could put the field of RNA translation initiation one step closer to not only understanding regulatory processes, but also assist in generating therapeutic protein products.
Dr. Wendy Gilbert earned her Bachelor of Arts degree in Molecular Biology at Princeton University before completing her PhD at the University of California, San Francisco under the supervision of Dr. Christine Guthrie. She then went on to complete her postdoctoral fellowship with Dr. Jennifer Doudna through the Howard Hughes Medical Institute and the University of California, Berkeley. Dr. Gilbert then joined as faculty at the Massachusetts Institute of Technology in the Department of Biology before receiving tenure in 2013. Now, Dr. Gilbert is an Associate Professor of Molecular Biophysics and Biochemistry at Yale University. She has mentored over 30 students and postdoctoral fellows, contributed to service and teaching, and recently received the Poorvu Price for Academic Innovation at Yale.