Author: Brooke Wolford
Editors: Jimmy Brancho, Shweta Ramdas, Belle Moyers
Think back to the last time you visited your primary care physician. Was the health care provider using a laptop or tablet to take notes and update your health information? In many doctors’ offices across the country your health records have gone digital. In addition to their exciting potential to help doctors’ offices reduce human error and better serve patients, electronic health records (EHRs) also make available a new source of “big data” for researchers.
EHRs are patient-specific digital records your health care provider maintains. The information in your EHR helps your doctor efficiently track your health over time and helps researchers learn more about diseases, which ultimately improves the clinical care your doctor provides to you and other patients. Believe it or not, EHRs from patients like you and me have already helped researchers make discoveries that improve health care for everyone!
How EHRs can be used to help researchers, clinicians, and patients
EHRs are a long-term, consolidated record of a person’s diagnoses, family history, past procedures, blood work, etc. One type of information that researchers identify from EHRs is a person’s disease status. Most frequently, diseases are systematically coded in EHRs through ICD codes (International Classification of Diseases).
The ICD codes are maintained by the World Health Organization and used worldwide for billing and insurance purposes, morbidity statistics, and now, research. These codes are used to classify diseases, often in a manner that is very specific to the causes, symptoms, or variations of the disease. For example, type 2 diabetes (T2D) can be described by over 20 ICD codes, which represent T2D with and without a variety of associated complications such as eye or circulatory problems. The codes are diverse, specific, and sometimes entertaining—the latest ICD code revision, ICD-10, includes codes such as Y93D9 which is “activity involving arts and handcrafts” and Z631 which is “problems in relationship with in-laws.”
Using ICD codes, researchers can use EHR data to improve medicine in a variety of ways. One possible application is to track patient prognosis—did changes in diet and exercise of a diabetic patient improve blood sugar levels and/or decrease T2D complications over time?
Other applications aim to find associations between diseases by asking: do certain diseases frequently occur together across multiple patients? Having such information can help clinicians diagnose and treat their patients better, in cases where a patient has a disease that is known to occur with another disease. A 2009 publication by Michigan scientists explored this question, and found the answer to be yes. The study did not only confirm known associations (e.g. T2D and obesity), but also discovered new associations. A heart condition called tricuspid valve regurgitation, which occurs when a heart valve does not work properly, was found to be associated with past use of tobacco. This novel association makes sense to clinicians who have long known that tobacco use increases the risk of heart disease.
EHRs + genetic data = Genome Wide Association Studies
Thanks to powerful computers, we can now make sense of the large and complex collections of health data from EHRs, a task which would be slow or impossible otherwise. Many large research consortiums have started linking EHRs and genetic data. Oftentimes a blood sample is obtained from a patient, and then DNA is extracted. The blood or DNA is saved in a biobank or biorepository and the anonymous genetic data is safely stored for access by researchers.
Here at the University of Michigan, the Michigan Genomics Initiative (MGI) allows patients at the U of M Health System to opt-in to help biomedical research (Figure 1). Individuals provide a blood sample that is analyzed to obtain genetic variant information at millions of single nucleotide polymorphisms (SNPs), which refer to a difference in a single nucleotide. The SNP data are linked to clinical data like patients’ EHRs and questionnaires. As of the time of this post, MGI has over 32,000 participants. Other examples of combining EHRs and biobank specimens are Vanderbilt University’s BioVU and Norway’s HUNT study which have over 150,000 and 80,000 blood samples respectively.
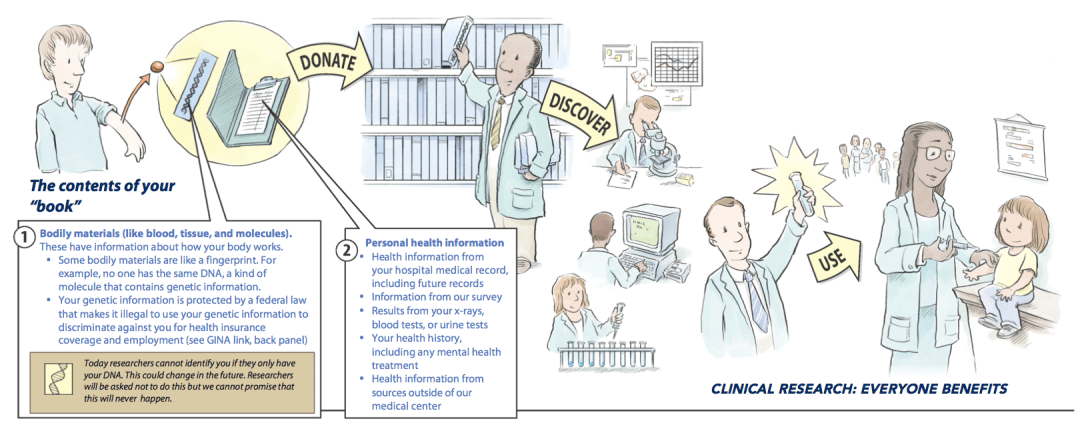
Figure 1. The Michigan Genomics Initiative includes this cartoon in their consent form. It walks potential MGI participants through (1) the collection of a blood sample and DNA and (2) the use of personal health information in the form of EHRs. The combination of this clinical and genetic data aids in clinical research, which has benefits for everyone.
In 2007 the NIH announced the Electronic Medical Records and Genomics (eMERGE) Network to encourage and fund the combination of EHRs and genetic data to revolutionize biomedical research. In 2015, under the leadership of Dr. Francis Collins, the NIH launched the Precision Medicine Initiative (PMI). Under PMI, researchers are working towards amassing a million-person cohort with clinical data from EHRs and genetic data. This diverse cohort is an exciting prospect for researchers because many current studies are limited to individuals of European ancestry, but we can learn more about the human genome, and better serve non-European patients, by looking at samples from all ethnicities. The PMI Cohort Program begins in 2017, so your doctor may ask you soon if you would like to enroll and contribute your EHR and genetic information to research!
The combination of EHRs and genetic data from blood samples in biobanks is a powerful tool for research. One way to harness both medical and genotype data (data on a person’s genetic makeup) is through a statistical method called genome-wide association studies (GWAS). Using EHRs, scientists can categorize a large group of people into cases, those with a certain disease, and controls, who are unaffected by the disease. By looking at many SNPs in the genome, scientists then can find genetic variants that are associated with the cases specifically, and rarely seen in the controls (Figure 2).
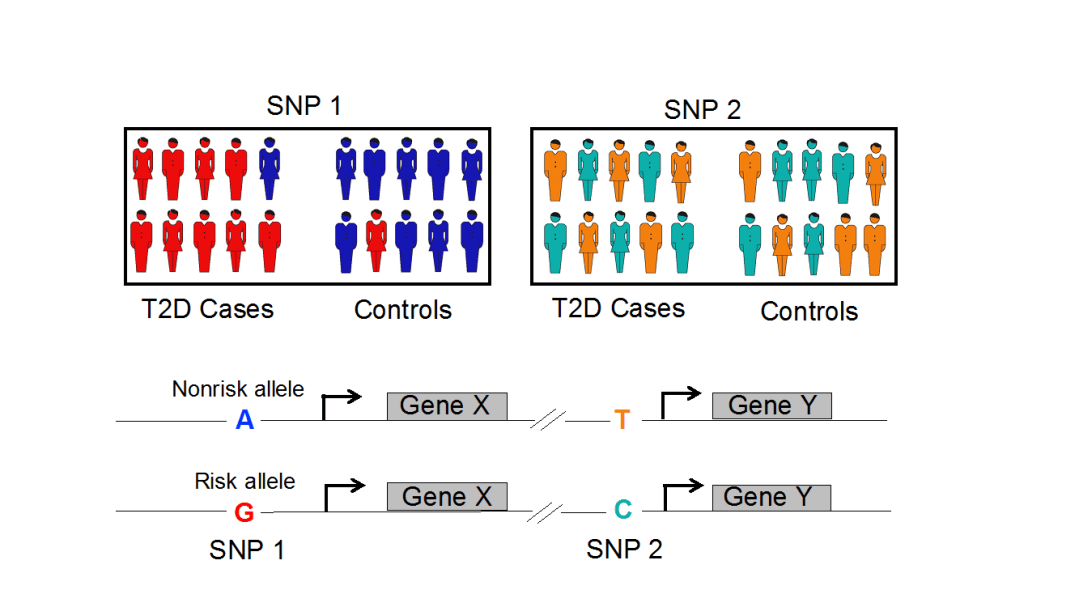
Figure 2. An illustration of a genome-wide association study (GWAS) for Type 2 diabetes. Here, the risk allele G (red) at SNP1 is seen in the majority of cases and rarely in controls. Therefore, this genotype is statistically associated with T2D. At SNP 2 neither genotype is seen in cases versus controls so the SNP is not associated with the disease.
By finding statistically significant associations between genotype and a specific medical condition, GWAS helps to pin-point the genetic variants that cause disease. For example, the MGI data allows researchers to find several SNPs in the genome that are seen in T2D cases but not in controls. When patients have these genetic variants and thus are at an increased risk for a disease like T2D, doctors can use this information to encourage lifestyle changes like diet and exercise in an effort to decrease the chance of developing the disease.
EHRs are revolutionizing the way scientists perform biomedical research in large patient populations. As scientists analyze more EHRs and genetic data their discoveries will continue to benefit patients like you and me. So, the next time you see a health care provider clicking away at a laptop screen, remember the many values of EHRs. Not only does it help your physician provide accurate and personalized care to you, but your EHR may one day help researchers better understand human disease, which helps everyone!
So what do EHRs have to do with Neanderthals? In Part III we will explore how EHRs and the Neanderthal genome allowed researchers to identify how Neanderthal DNA affects modern day human health. I explored the sequencing of the Neanderthal genome in “Interpreting ancient DNA: not so easy a caveman can do it” so please revisit Part I in preparation for Part III!
About the author
 Brooke is a PhD student in the Department of Computational Medicine and Bioinformatics. Her research focuses on understanding the genetic causes of complex diseases, specifically type 2 diabetes and cardiovascular traits. She is broadly interested in how genomics research can apply to clinical sequencing and patient care to fulfill the promise of Precision Medicine. Originally from North Carolina, Brooke holds a Bachelor of Science in Quantitative Biology from the University of North Carolina at Chapel Hill. When not staring at lines of code and sequence data, she enjoys reading, a good NETFLIX binge, and baking. Read more from Brooke here.
Brooke is a PhD student in the Department of Computational Medicine and Bioinformatics. Her research focuses on understanding the genetic causes of complex diseases, specifically type 2 diabetes and cardiovascular traits. She is broadly interested in how genomics research can apply to clinical sequencing and patient care to fulfill the promise of Precision Medicine. Originally from North Carolina, Brooke holds a Bachelor of Science in Quantitative Biology from the University of North Carolina at Chapel Hill. When not staring at lines of code and sequence data, she enjoys reading, a good NETFLIX binge, and baking. Read more from Brooke here.
Image Sources
Figure 1: https://www.michigangenomics.org/participate.html
Figure 2: Brooke Wolford

One thought on “How Your Electronic Health Records Could Help Biomedical Research”