Author: Jessica Cote
Editors: Zena Lapp, Christina Vallianatos, and Whit Froehlich
The Human Genome Project is one of the greatest scientific accomplishments in recent history— this international collaboration identified almost all of the ~20,500 genes in the human body, known collectively as the genome. Now that scientists know the details of these genes, they are better able to understand and treat human diseases associated with genetic factors. However, despite the immense effort put forth by over 30 research labs for 13 years (1990-2003), the information we gained from this project is limited. Genes serve as guidebooks for cells in the body to build proteins; genes themselves don’t perform the necessary cellular functions—proteins do. So, while scientists have now known the nitty-gritty of thousands of human genes for a while, the details of their protein products, known collectively as the proteome, are still quite puzzling.
Until about 40 years ago, most scientists believed the “one gene, one protein” hypothesis, which claims that each gene in our genome provides the instructions to build one single protein product. By this hypothesis, the human body should have about 20,500 proteins—but the actual human proteome may contain billions of proteins. So how can there be more proteins than genes? Recent research has debunked the “one gene, one protein” hypothesis, showing that the process by which genes “code” for proteins is much more complex than was previously thought. Here’s what we know now: due to a process called alternative splicing, one gene often produces two or more distinct proteins, or isoforms, which can serve unique (and sometimes opposing) functions. In one extreme example, a single gene involved in the development of the fruit fly brain may produce over 38,000 protein isoforms. These proteins help guide neurons to grow to their final destinations, and the many isoforms likely interact with different molecules in distinct areas of the brain.
How does alternative splicing work?
Let’s explore an analogy for alternative splicing. BioTrain, an imaginary company, is building two trains: one will carry passengers throughout a large city, and the other will transport farming equipment, animals, and a few passengers between farms. These trains represent protein isoforms, each one built to serve a special function in a particular location. The urban and rural regions represent different places in the body—like the heart and brain—that require different cellular activities to be performed by unique protein isoforms.
BioTrain Headquarters has sent a drawing to the urban and rural teams to show how many passenger and freight cars they are allowed to include in their trains (see Figure 1). HQ tells the teams to build only what they need because company resources are scarce. This original drawing from HQ represents a single gene, activated in cells in both the heart and the brain, which serves as a guide for the cells to build their proteins. Each car in the sketch represents an exon, which is the part of a gene that actually encodes a protein segment.
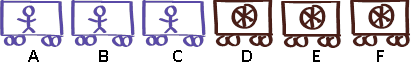
Each team is now using the original drawing from HQ as a guide to build its train. The teams’ processes of selecting specific train cars for their regions represents the alternative splicing mechanism, which—depending on various factors such as the location of the cell in an organ, or the developmental stage or sex of the organism—includes copies of some or all of the exons from the original gene in the final version of the guidebook to direct protein production.
The urban team designs a train with maximum passenger space, and no freight storage, to carry passengers throughout the city (see Figure 2.) This drawing represents a strand of messenger RNA, which transfers information from the original gene to the cellular machinery that builds proteins.

Because they need to transport farming equipment, animals, and some passengers between farms, the rural team decides to include all available freight cars and one passenger car in their train diagram (see Figure 3.) Similarly, this drawing represents a strand of messenger RNA that will build a different protein isoform, which differs greatly from the first.

Finally, the teams will use these new diagrams to build their trains, which represent two different protein isoforms that fulfill specific functions for cells in different organs. About 95% of all human multi-exon genes go through alternative splicing, so this system creates an enormous amount of biological diversity in our bodies. In this example, one gene guides production of two proteins through an alternative splicing method known as exon skipping. In other cases, a gene can guide the creation of tens to thousands of proteins through exon skipping, as well as other types of alternative splicing.
Why is the diversity of proteins produced by alternative splicing so important?
Actually, no one really knows. One popular theory is that protein diversity is a major contributor to organismal complexity, particularly for brain function. Our many alternative splices may explain how humans are capable of incredibly complicated behaviors such as communicating with spoken language, while other organisms—even those that are considered quite intelligent, like chimps—are not. Here’s some support for this theory: alternative splicing regulation varies widely among species, and the frequency at which it occurs increases with species complexity. For example, there are more instances of alternative splicing in humans than fruit flies. In addition, alternative splicing is particularly prevalent in the nervous system (including the brain) of vertebrates. Finally, mutations in proteins that are involved in the regulation of alternative splicing in the nervous system are associated with neurological disorders in humans, suggesting that normal splicing mechanisms are essential for our complex brains to function properly.
But there are weaknesses to the organismal complexity theory. There have been fewer studies done on fruit flies than humans, which may be skewing interpretations of comparisons between these organisms. It’s also unclear how much each protein type really contributes. Sure, there are more protein isoforms in humans than flies, but are these isoforms always adding value? In some cases, the function of one isoform appears identical to the function of the next, and the presence of both may not necessarily add to the complexity of the organism. Along with these uncertainties, other questions remain unanswered about alternative splicing and why it exists. For example, does alternative splicing serve another purpose, like allowing individuals of the same species to have unique traits?
These and other questions will keep researchers busy for many years as they work to understand the intricacies of alternative splicing. And while teams are busy investigating this protein diversity, the rest of us can learn about and celebrate diversity on a much larger scale: that of humans, whose complex behaviors and unique personalities constantly remind us how much more there is to learn about the microscopic details of our bodies.
About the author:
 Jess is a PhD student in the Neuroscience Program at the University of Michigan. She is studying how brain-specific forms of SH2B1, a gene associated with severe obesity in children, influence feeding behavior, energy use, and other aspects of metabolism. Before starting graduate school, Jess got her BS in Neuroscience with a minor in Hispanic Studies from Trinity College in Hartford, CT. She then moved to Chile to do neuroscience research with support from a Fulbright grant. In her free time, Jess enjoys playing tennis, doing yoga, going to concerts, and playing games with friends. Connect with her on LinkedIn.
Jess is a PhD student in the Neuroscience Program at the University of Michigan. She is studying how brain-specific forms of SH2B1, a gene associated with severe obesity in children, influence feeding behavior, energy use, and other aspects of metabolism. Before starting graduate school, Jess got her BS in Neuroscience with a minor in Hispanic Studies from Trinity College in Hartford, CT. She then moved to Chile to do neuroscience research with support from a Fulbright grant. In her free time, Jess enjoys playing tennis, doing yoga, going to concerts, and playing games with friends. Connect with her on LinkedIn.
Read all posts by Jess here.
